Chapter 16: The Molecular basis of inheritance
16.1: DNA is the genetic material
The Search for the Genetic Material: Scientific Inquiry
- T.H. Morgan: worked with Drosophila melanogaster and determined gene's exist as parts of chromosomes.
-Chromosomes are made up of DNA and protein
- it was not established which of the two represented hereditary material at first
-more was known about proteins than nucleic acids, and based on this knowledge, proteins
were considered a better candidate prior to research to prove different.
Evidence That DNA Can Transform Bacteria
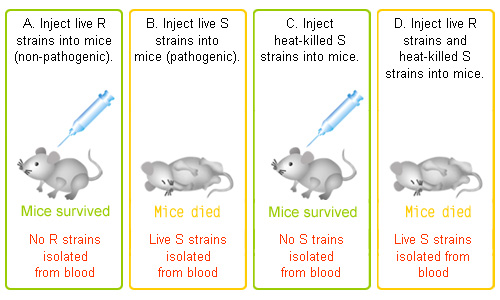
-Griffith: While developing a vaccine for pneumonia, and studying Streptococcus pneumoniae, the bacteria that causes pneumonia in mammals, Griffith discovered the transformation of bacteria. (see experiment below)
-transformation: a change in genotype and phenotype due to the assimilation of external DNA by a cell
The Search for the Genetic Material: Scientific Inquiry
- T.H. Morgan: worked with Drosophila melanogaster and determined gene's exist as parts of chromosomes.
-Chromosomes are made up of DNA and protein
- it was not established which of the two represented hereditary material at first
-more was known about proteins than nucleic acids, and based on this knowledge, proteins
were considered a better candidate prior to research to prove different.
Evidence That DNA Can Transform Bacteria
-Griffith: While developing a vaccine for pneumonia, and studying Streptococcus pneumoniae, the bacteria that causes pneumonia in mammals, Griffith discovered the transformation of bacteria. (see experiment below)
-transformation: a change in genotype and phenotype due to the assimilation of external DNA by a cell
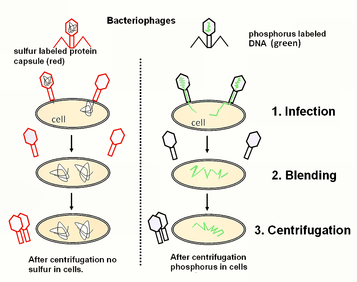
-Hershey and Chase: proved that DNA was the genetic material using T2 phage, showing that only DNA entered the E. coli during infection. (see experiment below using sulfur tagged protein and phosphorous tagged DNA)
-virus: a little more than DNA (or sometimes RNA) enclosed by a protein capsule
- to produce more viruses a virus must infect a cell and take over the cell's metabolic machinery.
-Phage/ bacteriophage: a virus that infects and replicates within a bacterium. The term is derived from "bacteria" and the "to devour."
-virus: a little more than DNA (or sometimes RNA) enclosed by a protein capsule
- to produce more viruses a virus must infect a cell and take over the cell's metabolic machinery.
-Phage/ bacteriophage: a virus that infects and replicates within a bacterium. The term is derived from "bacteria" and the "to devour."
Additional Evidence that DNA is the Genetic Material
-Chargaff: After analyzing the (nitrogenous) base composition of different organisms, he discovered 2 important findings that became Chargaff's Rules:
1) The composition of DNA varied between species.
2) The ratios of Adenine and Thymine in DNA were relatively the same and that the ratios of Cytosine and Guanine were relatively the same (it was later proven with updated techniques that these bases exist in exact 1:1 ratios with eachother).
Building a Structural Model of DNA
-DNA was established as the genetic material, now scientists needed to find its structure to account for its role as the hereditary material.
-Watson and Crick: Watson joined Crick with a x-ray crystallograph provided by Rosalind Franklin
- Watson was familiar with the pattern of helical molecules and recognized this pattern in the image of the DNA.
- The pattern also implied a double-stranded nature of the DNA molecule.
-This led to the naming of the double helix
-Franklin: determined the sugar and phosphate groups were on the outside of the molecule due to the hydrophobic nature of nitrogenous bases and the negatively charge phosphate groups (that wouldn't fair well positioned against eachother because of their charges repelling).
- Using this Warson and Crick came up with an antiparallel model for the backbones, meaning their subunits run in opposite directions.
- Franklin also determined that the helix made a full turn every 3.4 nm, with bases stacked 0.34 nm apart, making it so there are
10 base pairs per turn in a DNA molecule.
-Though Chargaff had determined the ratios of A, T, C and G, Watson an Crick determined that because of the constant width of the double helix, a purine and a pyrimidine would have to pair in order for the width of the molecule to remain constant. Therefore A paired with T and C paired with G from Chargaff's provided ratios.
-Watson and Crick published their proposal for DNA structure in 1953, and it remains to this day.
-Though the bases pair specifically, there is not specification to the linear ordering of bases along a single strand, and there can be countless variations, accounting for the unique base sequence of each gene.
16.2: Many Proteins work together in DNA Replication and Repair
-The discovered structure of DNA, and its specific base pairing, led to a proposed copying mechanism for the molecule.
The Basic Principle: Base Pairing to a Template Strand
-Specific base pairing allows for determining one strand's sequence by just referring to the other strand.
- The two strands are complimentary, meaning that each strand has the information necessary to reconstruct the other.
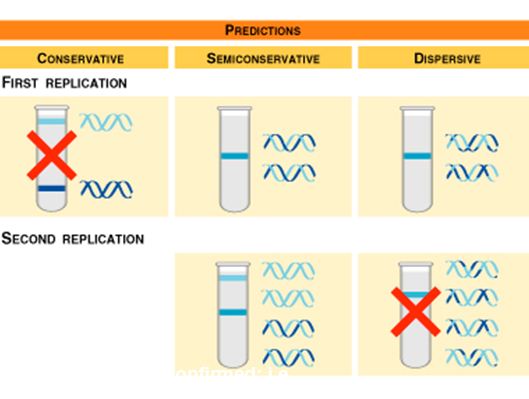
-Semiconservative Model of Replication: Each strand serves as a template for ordering nucleotides into a new, complementary strand. There are two identical DNA molecules at the end of the process, each containing one original strand from the parent molecule.
- The conservative model, where the parental strand is maintained, and the dispersive model, where each molecule is a mixture of parent and newly formed, were both disproven thanks to Meselson and Stahl. (see experiment below)
-Chargaff: After analyzing the (nitrogenous) base composition of different organisms, he discovered 2 important findings that became Chargaff's Rules:
1) The composition of DNA varied between species.
2) The ratios of Adenine and Thymine in DNA were relatively the same and that the ratios of Cytosine and Guanine were relatively the same (it was later proven with updated techniques that these bases exist in exact 1:1 ratios with eachother).
Building a Structural Model of DNA
-DNA was established as the genetic material, now scientists needed to find its structure to account for its role as the hereditary material.
-Watson and Crick: Watson joined Crick with a x-ray crystallograph provided by Rosalind Franklin
- Watson was familiar with the pattern of helical molecules and recognized this pattern in the image of the DNA.
- The pattern also implied a double-stranded nature of the DNA molecule.
-This led to the naming of the double helix
-Franklin: determined the sugar and phosphate groups were on the outside of the molecule due to the hydrophobic nature of nitrogenous bases and the negatively charge phosphate groups (that wouldn't fair well positioned against eachother because of their charges repelling).
- Using this Warson and Crick came up with an antiparallel model for the backbones, meaning their subunits run in opposite directions.
- Franklin also determined that the helix made a full turn every 3.4 nm, with bases stacked 0.34 nm apart, making it so there are
10 base pairs per turn in a DNA molecule.
-Though Chargaff had determined the ratios of A, T, C and G, Watson an Crick determined that because of the constant width of the double helix, a purine and a pyrimidine would have to pair in order for the width of the molecule to remain constant. Therefore A paired with T and C paired with G from Chargaff's provided ratios.
-Watson and Crick published their proposal for DNA structure in 1953, and it remains to this day.
-Though the bases pair specifically, there is not specification to the linear ordering of bases along a single strand, and there can be countless variations, accounting for the unique base sequence of each gene.
16.2: Many Proteins work together in DNA Replication and Repair
-The discovered structure of DNA, and its specific base pairing, led to a proposed copying mechanism for the molecule.
The Basic Principle: Base Pairing to a Template Strand
-Specific base pairing allows for determining one strand's sequence by just referring to the other strand.
- The two strands are complimentary, meaning that each strand has the information necessary to reconstruct the other.
-Semiconservative Model of Replication: Each strand serves as a template for ordering nucleotides into a new, complementary strand. There are two identical DNA molecules at the end of the process, each containing one original strand from the parent molecule.
- The conservative model, where the parental strand is maintained, and the dispersive model, where each molecule is a mixture of parent and newly formed, were both disproven thanks to Meselson and Stahl. (see experiment below)
DNA Replication: a Closer Look
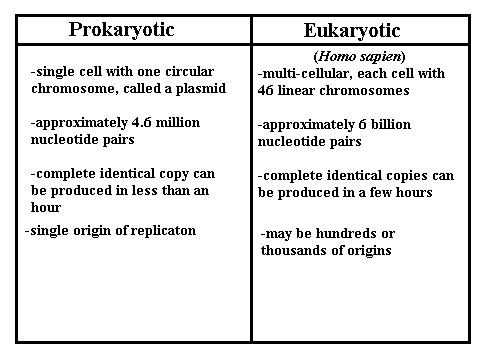
-The model of prokaryotic replication is more well-known, however, what we know about eukaryotic replication suggest that the processes are fundamentally similar.
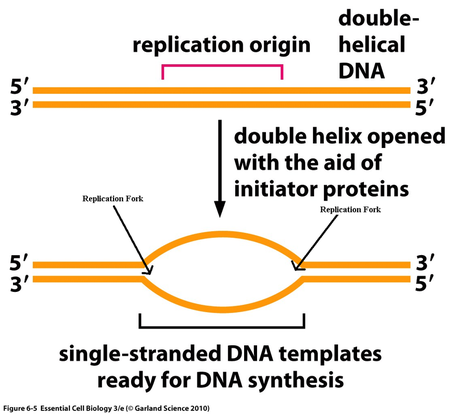
-origin/s of replication: short stretches of DNA with a specific sequence of nucleotides that is recognized by proteins that initiate replication.
-Proteins that initiate replication recognize and bind the specific origin/s, separating the two strands and opening a replication "bubble/s," forming replication forks at both ends of the opening.
-origin/s of replication: short stretches of DNA with a specific sequence of nucleotides that is recognized by proteins that initiate replication.
-Proteins that initiate replication recognize and bind the specific origin/s, separating the two strands and opening a replication "bubble/s," forming replication forks at both ends of the opening.
-Replication proceeds in both directions of the forks, with the aid of unwinding proteins:
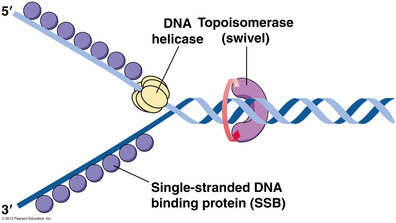
-Helicases: unwind the double helix at the replication forks, separating the strands to make them available as templates for continued replication
-Single-strand binding proteins: after the strands are separated, these proteins bind to the individual strands to prevent
re-pairing of bases on the parental strands.
-Topoisomerase: helps relieve strain ahead of the replication forks caused by the helicase untwisting at the forks.
-Helicases: unwind the double helix at the replication forks, separating the strands to make them available as templates for continued replication
-Single-strand binding proteins: after the strands are separated, these proteins bind to the individual strands to prevent
re-pairing of bases on the parental strands.
-Topoisomerase: helps relieve strain ahead of the replication forks caused by the helicase untwisting at the forks.
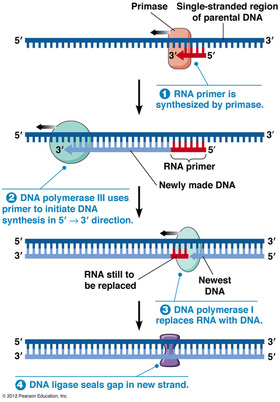
-Enzymes that synthesize DNA cannot initiate the synthesis of a polynucleotide, they require an RNA primer (generally 5-10 nucleotides long), synthesized by the enzyme primase. Once the short stretch of RNA is in place, enzymes called DNA polymerases begin to add nucleotides to the end of the existing chain.
-DNA Polymerase I: replaces RNA nucleotides of the primer with DNA nucleotides
-DNA Polymerase III: adds DNA nucleotides to the existing RNA primer
-DNA Polymerase I: replaces RNA nucleotides of the primer with DNA nucleotides
-DNA Polymerase III: adds DNA nucleotides to the existing RNA primer
- Like ATP, each nucleotide added to the growing strand consists of 3 phosphate groups, and are chemically reactive, and lose two of these phosphate groups upon their addition.
- this serves as a coupled exergonic reaction, which fuels polymerization
Antiparallel Elongation
-The two strands of a DNA double helix are antiparallel, or oriented in opposite directions. Therefore, during replication, the two newly formed strands must be antiparallel to their parental strand.
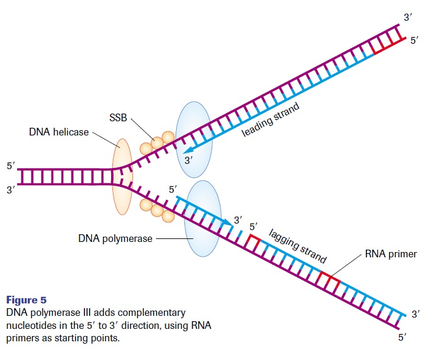
-DNA polymerases can only add nucleotides to the 3' end of a growing polynucleotide. A new DNA strand can only be formed in a 5' to 3' direction. This results in the leading and lagging strands during DNA synthesis. (refer to text image page 323)
-leading strand: continuously synthesized in the direction of the replication fork.
-requires a single primer
-lagging strand: synthesized in fragments, called Okazaki fragments, in the opposite direction of the replication fork.
-requires multiple primers, each fragment must be primed separately.
-DNA Polymerase I replaces these primers as synthesis continues, by using the 3' end of the existing fragment.
-DNA ligase joins the DNA replacement with the following fragment
- this serves as a coupled exergonic reaction, which fuels polymerization
Antiparallel Elongation
-The two strands of a DNA double helix are antiparallel, or oriented in opposite directions. Therefore, during replication, the two newly formed strands must be antiparallel to their parental strand.
-DNA polymerases can only add nucleotides to the 3' end of a growing polynucleotide. A new DNA strand can only be formed in a 5' to 3' direction. This results in the leading and lagging strands during DNA synthesis. (refer to text image page 323)
-leading strand: continuously synthesized in the direction of the replication fork.
-requires a single primer
-lagging strand: synthesized in fragments, called Okazaki fragments, in the opposite direction of the replication fork.
-requires multiple primers, each fragment must be primed separately.
-DNA Polymerase I replaces these primers as synthesis continues, by using the 3' end of the existing fragment.
-DNA ligase joins the DNA replacement with the following fragment
Proofreading and Repairing DNA
-Initial pairing errors occur at a rate of one in 10^5 nucleotides.
-However, errors in a completed DNA molecule amount to only one in 10^10 nucleotides.
-This is because DNA polymerases proofread each nucleotide against its template as soon as it is covalently bonded to the growing strand.
-Upon finding an incorrect nucleotide, the polymerase will remove it and continue synthesis. However, sometimes errors elude polymerases.
-Mismatch repair: other enzymes remove and replace incorrect nucleotides missed by DNA Pol.
-There are almost 100 repair enzymes in prokaryotes and about 130 have been identified in eukaryotes, suggesting evolution of repair enzymes over time.
-in a particularly aggressive form of colon cancer, there is a hereditary defect in one of these repair enzymes, which allows cancer-causing errors to accumulate faster than what is normal.
-Incorrectly paired or altered nucleotides are not only caused by errors in replication, they can occur spontaneously or after exposure to harmful chemical or physical agents, like X-rays.
-However, these changes are usually corrected before they become permanent mutations, which perpetuate through successive replications.
-Mutation: permanent in the DNA sequence
-nuclease: a DNA-cutting enzyme, that excises the segment of the DNA strand containing the mismatch error, and the gap is then filled with the correct nucleotides by DNA polymerase, using the undamaged strand as a template, and attached by DNA ligase.
-nucleotide exicision repair (NER): DNA repair system that cuts out incorrect segments and replaces them with correct nucleotides.
Evolutionary Significance of Altered DNA Nucleotides
-The faithful copying of the genome and reparation of damage is paramount to every organism.
-mutations can change the phenotype of an organism.
-If (and ONLY if) these mutations occur in germ cells, which give rise to gametes, mutations will be passed on to successive generations.
-The vast majority have no effect or are harmful, though a very small percentage can be beneficial.
-Mutations are the original source of phenotypic variation on which natural selection operates.
Replicating Ends of DNA Molecules
-For linear DNA, as found in eukaryotes, DNA polymerase cannot synthesize the 5' ends of daughter DNA strands because it can only add nucleotides to an existing 3' end.
-As a result of repeated replications, the DNA molecules that result have staggered ends, and become shorter and shorter.
-Prokaryotes do not have this problem because of the circular shape of their DNA.
-What protects the genes of the eukaryotic chromosomes are special nucleotide sequences called telomeres.
-Telomeres do not contain genes, but consist of multiple repetitions of one short nucleotide sequence.
-specific proteins associated with telomeric DNA prevent the staggered ends of the daughter molecule from activating the cell's system for monitoring DNA damage.
-Telomeric DNA acts as a buffer zone that provides postponement to the erosion of genetic material.
-It has been proposed that the shortening of telomeres is somehow connected to the aging process of certain tissues and even to the aging of an organism as a whole.
-Germ cells cannot shorten over time or the offspring of that organism would be at risk for not receiving all essential genes.
-they have an enzyme called telomerase that catalyzes the lengthening of their telomeres.
-this enzyme is usually inactive in somatic cells.
-certain cancerous cells activate the synthesis of this enzyme, allowing for infinite divisions.
-researchers are focused on the inactivation of telomerase for certain somatic cell cancers.
16.3: A Chromosome Consists of a DNA Molecule Packed Together with Proteins
-In the cell, eukaryotic DNA is precisely combined with a large amount of protein.
-Chromatin: the complex of DNA and protein, packaging that allows for the DNA to fit in the nucleus
-proteins called histones are responsible for the primary level of DNA packaging. These proteins have been highly conserved throughout evolution, which maintains their importance in the organization of DNA.
-The DNA is wound around the histones, forming beads, called nucleosomes.
-Chromatin undergoes many changes in respect to its packing during the cell cycle.
-Heterochromatin: highly condensed state of chromatin during interphase, chromosomes in this form are less accessible for transcription
-Euchromatin: loosely packed state of chromatin while cell is not dividing, allows access by transcription machinery
-Initial pairing errors occur at a rate of one in 10^5 nucleotides.
-However, errors in a completed DNA molecule amount to only one in 10^10 nucleotides.
-This is because DNA polymerases proofread each nucleotide against its template as soon as it is covalently bonded to the growing strand.
-Upon finding an incorrect nucleotide, the polymerase will remove it and continue synthesis. However, sometimes errors elude polymerases.
-Mismatch repair: other enzymes remove and replace incorrect nucleotides missed by DNA Pol.
-There are almost 100 repair enzymes in prokaryotes and about 130 have been identified in eukaryotes, suggesting evolution of repair enzymes over time.
-in a particularly aggressive form of colon cancer, there is a hereditary defect in one of these repair enzymes, which allows cancer-causing errors to accumulate faster than what is normal.
-Incorrectly paired or altered nucleotides are not only caused by errors in replication, they can occur spontaneously or after exposure to harmful chemical or physical agents, like X-rays.
-However, these changes are usually corrected before they become permanent mutations, which perpetuate through successive replications.
-Mutation: permanent in the DNA sequence
-nuclease: a DNA-cutting enzyme, that excises the segment of the DNA strand containing the mismatch error, and the gap is then filled with the correct nucleotides by DNA polymerase, using the undamaged strand as a template, and attached by DNA ligase.
-nucleotide exicision repair (NER): DNA repair system that cuts out incorrect segments and replaces them with correct nucleotides.
Evolutionary Significance of Altered DNA Nucleotides
-The faithful copying of the genome and reparation of damage is paramount to every organism.
-mutations can change the phenotype of an organism.
-If (and ONLY if) these mutations occur in germ cells, which give rise to gametes, mutations will be passed on to successive generations.
-The vast majority have no effect or are harmful, though a very small percentage can be beneficial.
-Mutations are the original source of phenotypic variation on which natural selection operates.
Replicating Ends of DNA Molecules
-For linear DNA, as found in eukaryotes, DNA polymerase cannot synthesize the 5' ends of daughter DNA strands because it can only add nucleotides to an existing 3' end.
-As a result of repeated replications, the DNA molecules that result have staggered ends, and become shorter and shorter.
-Prokaryotes do not have this problem because of the circular shape of their DNA.
-What protects the genes of the eukaryotic chromosomes are special nucleotide sequences called telomeres.
-Telomeres do not contain genes, but consist of multiple repetitions of one short nucleotide sequence.
-specific proteins associated with telomeric DNA prevent the staggered ends of the daughter molecule from activating the cell's system for monitoring DNA damage.
-Telomeric DNA acts as a buffer zone that provides postponement to the erosion of genetic material.
-It has been proposed that the shortening of telomeres is somehow connected to the aging process of certain tissues and even to the aging of an organism as a whole.
-Germ cells cannot shorten over time or the offspring of that organism would be at risk for not receiving all essential genes.
-they have an enzyme called telomerase that catalyzes the lengthening of their telomeres.
-this enzyme is usually inactive in somatic cells.
-certain cancerous cells activate the synthesis of this enzyme, allowing for infinite divisions.
-researchers are focused on the inactivation of telomerase for certain somatic cell cancers.
16.3: A Chromosome Consists of a DNA Molecule Packed Together with Proteins
-In the cell, eukaryotic DNA is precisely combined with a large amount of protein.
-Chromatin: the complex of DNA and protein, packaging that allows for the DNA to fit in the nucleus
-proteins called histones are responsible for the primary level of DNA packaging. These proteins have been highly conserved throughout evolution, which maintains their importance in the organization of DNA.
-The DNA is wound around the histones, forming beads, called nucleosomes.
-Chromatin undergoes many changes in respect to its packing during the cell cycle.
-Heterochromatin: highly condensed state of chromatin during interphase, chromosomes in this form are less accessible for transcription
-Euchromatin: loosely packed state of chromatin while cell is not dividing, allows access by transcription machinery