Chapter 18: Regulation of Gene expression
Bacteria Often Respond to Environmental Change by Regulating Transcription
-Bacteria that express only the genes whose products are needed by the cell conserve resources and energy, causing these bacteria to be favored by natural selection. They use two forms of metabolic control to achieve this:
1) feedback inhibition: cells can adjust the activity of the enzymes already present, the activity of the first enzyme in a pathway is inhibited by the pathway's end product
2) cells can also adjust the production level of certain enzymes by regulating the genes that code for the enzyme, and control occurs at the transcription level (genes are turned on or off by changes in the metabolic status of the cell)
- operon model
Operons: The Basic Concept
-Operon: an operator, a promoter, and the genes they control.
-A group of genes of related function exist in a single transcription unit, where a single promoter serves all the genes in the unit. Therefore, the single unit gives rise to multiple protein products of the functionally related genes in one transcript.
- this is beneficial because a single on-off switch, called an operator, can control the whole cluster of functionally related genes.
- the operator is located within the promoter (or sometimes between the promoter and the enzyme coding genes), and controls the access of RNA polymerase to the genes.
-Repressor: a protein that binds the operator, turning the operon off by preventing the binding of RNA polymerase, and therefore, transcription.
- repressors are the protein products of regulatory genes, which are controlled by their own promoters, and typically expressed continuously at low rates.
- Though repressors may be synthesized continuously, operators exist in two states, alternating between being bound to a repressor and not being bound to a repressor, the frequency of being bound to a repressor increases with the concentration of the repressor.
-Repressors are also allosteric inhibitors, meaning they have both an active and inactive form.
-active when the product of the operon is present and bound at an allosteric site, altering the active site of the protein, and giving it an increased affinity for the operator- repressing transcription.
-inactive when the product of the operon is not bound to the allosteric site, reducing its affinity for the operator.
-Bacteria that express only the genes whose products are needed by the cell conserve resources and energy, causing these bacteria to be favored by natural selection. They use two forms of metabolic control to achieve this:
1) feedback inhibition: cells can adjust the activity of the enzymes already present, the activity of the first enzyme in a pathway is inhibited by the pathway's end product
2) cells can also adjust the production level of certain enzymes by regulating the genes that code for the enzyme, and control occurs at the transcription level (genes are turned on or off by changes in the metabolic status of the cell)
- operon model
Operons: The Basic Concept
-Operon: an operator, a promoter, and the genes they control.
-A group of genes of related function exist in a single transcription unit, where a single promoter serves all the genes in the unit. Therefore, the single unit gives rise to multiple protein products of the functionally related genes in one transcript.
- this is beneficial because a single on-off switch, called an operator, can control the whole cluster of functionally related genes.
- the operator is located within the promoter (or sometimes between the promoter and the enzyme coding genes), and controls the access of RNA polymerase to the genes.
-Repressor: a protein that binds the operator, turning the operon off by preventing the binding of RNA polymerase, and therefore, transcription.
- repressors are the protein products of regulatory genes, which are controlled by their own promoters, and typically expressed continuously at low rates.
- Though repressors may be synthesized continuously, operators exist in two states, alternating between being bound to a repressor and not being bound to a repressor, the frequency of being bound to a repressor increases with the concentration of the repressor.
-Repressors are also allosteric inhibitors, meaning they have both an active and inactive form.
-active when the product of the operon is present and bound at an allosteric site, altering the active site of the protein, and giving it an increased affinity for the operator- repressing transcription.
-inactive when the product of the operon is not bound to the allosteric site, reducing its affinity for the operator.
Repressible and Inducible Operons: Two Types of Negative Gene Regulation
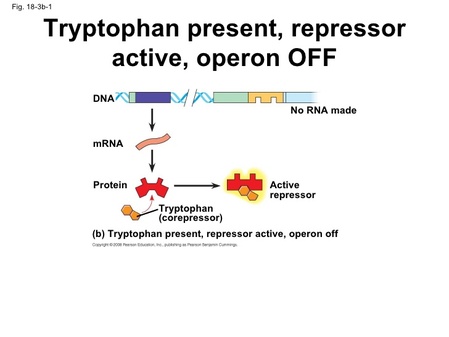
- Repressible operon (discussed above): usually expressed (on), but can be inhibited (repressed) if a specific small molecule bind allosterically to the repressor protein, increasing the repressors affinity for the operator.
-trp operon
- Repressible operon (discussed above): usually expressed (on), but can be inhibited (repressed) if a specific small molecule bind allosterically to the repressor protein, increasing the repressors affinity for the operator.
-trp operon
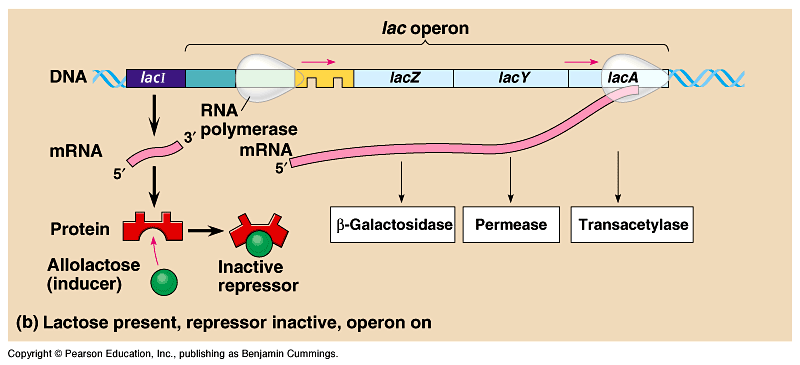
-Inducible operon: usually not expressed (off), but can be stimulated, when a specific small molecule interacts with a regulatory protein, decreasing the repressors affinity for the operator.
- lac operon: lactose is hydrolyzed into glucose and galactose, catalyzed by the enzyme B- galactosidase.
-few copies of this enzyme are present in an E. coli cell in the absence of lactose
-however, if lactose is introduced, the number of B-galactosidase molecules can increase a thousand-fold
within 15 minutes.
-The gene that codes for this enzyme is called lacZ, and is part of the lac operon, which includes two other genes
that code for enzymes that function in the use of lactose.
-The regulatory gene that codes for the repressor is called lacI, which switches off the gene by binding the operator.
- the lac repressor (unlike the trp repressor) is active by itself, and becomes inactive when it is bound to an inducer molecule, allolactose.
-This allows E. coli to save energy and resources in the absence of lactose, but to utilize lactose when it is present and glucose is scarce or absent.
- lac operon: lactose is hydrolyzed into glucose and galactose, catalyzed by the enzyme B- galactosidase.
-few copies of this enzyme are present in an E. coli cell in the absence of lactose
-however, if lactose is introduced, the number of B-galactosidase molecules can increase a thousand-fold
within 15 minutes.
-The gene that codes for this enzyme is called lacZ, and is part of the lac operon, which includes two other genes
that code for enzymes that function in the use of lactose.
-The regulatory gene that codes for the repressor is called lacI, which switches off the gene by binding the operator.
- the lac repressor (unlike the trp repressor) is active by itself, and becomes inactive when it is bound to an inducer molecule, allolactose.
-This allows E. coli to save energy and resources in the absence of lactose, but to utilize lactose when it is present and glucose is scarce or absent.
Inducible enzymes usually function in catabolic pathways, by producing enzymes only when the specific nutrient is available for breakdown.
Repressible enzymes usually function in anabolic pathways, synthesizing products when needed, but suspending production when the product is present in sufficient quantities.
Negative control: operons switched off by active form of the repressor only, acts to control/regulate the transcription of a gene based on need.
- both the trp and lac repressors are forms of negative control, though their repressors are activated and inactivated in opposite ways.
Positive control: regulatory protein directly interacts with genome to switch transcription on or off, acts to kick transcription into overdrive when called for.
Positive Gene Regulation
-When glucose and lactose are both present, E. coli prefers glucose and enzymes for glycolysis are continually present.
-Only when lactose is present and glucose is in short supply does E. coli synthesize appreciable quantities of enzymes for lactose breakdown.
- cyclic AMP (cAMP) accumulates in the absence of glucose and binds catabolite activator protein (CAP)
- CAP is an activator, a protein that binds DNA to stimulate the transcription of a gene.
-When cAMP binds CAP, CAP can bind to a specific sequence of DNA in the upstream end of the lac promoter, increasing the affinity of RNA polymerase to the promoter, resulting in an increased rate of transcription.
-Positive regulation because the attachment of CAP to the promoter, directly stimulates gene expression.
-If glucose becomes present again, the cAMP concentration falls, and CAP detaches from the operator.
-The lac operon is under dual control, negative by the lac repressor and positive by CAP.
- the state of the repressor determines whether the operon is transcribed at all
- the state of CAP determines the rate of transcription if the operator is not being repressed.
18.2: Eukaryotic Gene Expression is Regulated at Many Stages
Differential Gene Expression
-Though most cells in a multicellular organism contain the same genome, different genes are expressed by each cell type, allowing each cell to be specialized for its particular function
-Differential gene expression: the expression of different genes, by cells containing the same genome
-Gene expression is most commonly controlled at the transcription level by receiving signals from outside the cell, though there are other control methods.
-This is why, generally, gene expression is considered a gene being actively transcribed
Regulation of Chromatin Structure
-Chromatin is the complex formed between DNA and the histone proteins that package it, the basic unit, being a nucleosome
-This protein/DNA interaction not only allows for the chromosomes to fit into the nucleus, but also has a regulatory function in gene expression.
- location of the promoter with respect to the nucleosome and DNA attachment sites to the scaffold affect a genes ability to
be transcribed
-genes within highly condensed heterochromatin are usually not expressed.
-chemical modifications to the histone proteins and DNA can influence chromatin structure and gene expression
Histone Modifications and DNA Methylation
-Chemical modifications to histones play a role in gene regulation
-N terminus of each histone protrudes from nucleosome, making them accessible to various modifying enzymes that catalyze
the addition or removal of chemical groups:
-Histone acetylation (the addition of an acetyl group), generally promotes transcription by opening the chromatin structure.
-Histone methylation (the addition of a methyl group), leads to reduced transcription by further condensing chromatin.
-DNA methylation (the addition of a methyl group to DNA itself, usually a cytosine base), methylated genes are not expressed.
-These patterns of methylation are propogated through successive cellular divisions, keeping the same genes
silent in each new cell derived from the original.
-DNA methylation is important during embryonic development.
- development of specialized tissues (different types of cells express a different set of genes to become specialized for their particular function in a multicellular organism)
-Genomic imprinting: for certain genes, only one copy of the gene (maternal or paternal) is expressed (only ~9 chromosomes are known to have regions where impriting occurs)
-These chemical changes are all reversible, and do not change the DNA (therefore, they are not mutations) though they may remain throughout an organism's lifetime.
Epigenetic Inheritance
-Epigenetic inheritance: inheritance of traits transmitted by mechanisms that do not involve the nucleotide sequence, itself.
-DNA methylation patterns of an individual are largely erased and reestablished during gamete formation.
-Epigenetic variations (i.e. variations in DNA methylation patterns) are a proposed reason for the occurrence of genetically based diseases in idividuals with identical genomes (identical twins).
-Alterations in normal DNA methylation patterns are also found in some cancers, where they are associated with inappropriate gene expression.
Regulation of Transcription Initiation
- The regulation of of transcription initiation in bacteria and eukaryotes involves proteins that bind to DNA and either facilitate or inhibit binding of RNA polymerase.
Organization of a Typical Eukaryotic Gene
-Control elements: most eukaryotic genes have multiple segments of non-coding DNA that serves as binding sites for the proteins called transcription factors, which in turn regulate transcription.
- This is also important for the precise regulation of gene expression in different cell types.
The Roles of Transcription Factors
-Eukaryotic RNA polymerase requires the presence of transcription factors for initiation
-General transcription factors (GTFs) are proteins that are required for the transcription of all protein-coding genes.
- the interaction of GTFs and RNA pol. II with a promoter result in a low rate of initiation.
-Specific transcription factors (STFs) are proteins that are required to enhance the transcription rate of specific
genes.
Enhancers and Specific Transcription Factors
-Proximal control elements: control elements located close to the promoter.
-Distal control elements/ Enhancers: control elements located upstream or downstream from a gene, or sometimes within an intron. These elements are thought to bring about protein-mediated bending of the DNA, which bring about and increase in transcription.
-Genes can have multiple enhancers (which may not all be active simultaneously),
but each enhancer is generally associated with only one gene.
-Binding of specific transcription factors, either activators and repressors, to the control elements of enhancers can strongly increase or decrease the rate of transcription.
-Eukaryotic activator proteins have been found to share the presence of 2 domains within their structures,
a DNA binding domain and an activation domain which bind other regulatory proteins or components of the transcription machinery.
-Repressor proteins can inhibit gene expression by
- binding directly to a control element in the DNA, which blocks binding of activator
-Interfering with the activator itself, so it cannot bind the DNA
-Activators and repressors can also alter the structure of the chromatin, leading to increased or decreased transcription.
Combinatorial Control of Gene Activation
-