Chapter 17: Gene Expression: From Gene to protein
The Flow of Genetic Information
-Inherited traits are determined by genes, and the information content of genes is in the form of specific nucleotide sequencing along DNA strands.
-The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and RNA molecules involved in protein synthesis.
-Proteins are the link between genotype and phenotype
-Gene expression: The process by which DNA directs the synthesis of proteins ( or in some cases, just RNAs)
17.1 Genes Specify Proteins via Transcription and Translation
Evidence from the Study of Metabolic Defects:
-Garrod: genes dictate phenotypes through enzymes that catalyze specific reactions in the cell, symptoms of an inherited disease reflect a person's inability to make a particular enzyme.
-One gene-one enzyme hypothesis: a gene dictates the production of a specific enzyme
-cells synthesize and degrade most organic molecules via metabolic pathways, in which each chemical reaction in a sequence is catalyzed by a specific enzyme.
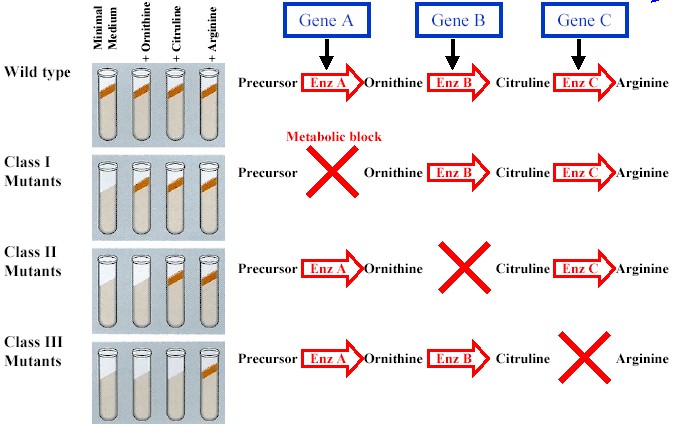
Nutritional Mutants in Neurospora
-Beadle and Tatum: mutations cause the synthesis of an enzyme to be blocked at a particular step in the production pathway due to the inability of the mutant to produce the enzyme needed to catalyze that particular step. (See experiment below)
-Inherited traits are determined by genes, and the information content of genes is in the form of specific nucleotide sequencing along DNA strands.
-The DNA inherited by an organism leads to specific traits by dictating the synthesis of proteins and RNA molecules involved in protein synthesis.
-Proteins are the link between genotype and phenotype
-Gene expression: The process by which DNA directs the synthesis of proteins ( or in some cases, just RNAs)
17.1 Genes Specify Proteins via Transcription and Translation
Evidence from the Study of Metabolic Defects:
-Garrod: genes dictate phenotypes through enzymes that catalyze specific reactions in the cell, symptoms of an inherited disease reflect a person's inability to make a particular enzyme.
-One gene-one enzyme hypothesis: a gene dictates the production of a specific enzyme
-cells synthesize and degrade most organic molecules via metabolic pathways, in which each chemical reaction in a sequence is catalyzed by a specific enzyme.
Nutritional Mutants in Neurospora
-Beadle and Tatum: mutations cause the synthesis of an enzyme to be blocked at a particular step in the production pathway due to the inability of the mutant to produce the enzyme needed to catalyze that particular step. (See experiment below)
The Products of Gene Expression
-Not all proteins are enzymes, hypothesis reformatted to one gene-one protein
-Many proteins are composed of two or more polypeptide chains, hypothesis reformatted to one gene- one polypeptide
-Still not completely accurate since alternative splicing allows for a single gene to code for multiple polypeptides and some genes solely code for RNA and never become polypeptides.
Basic Principles of Transcription and Translation
-Genes code for proteins, but are not directly responsible for their synthesis. RNA bridges the gap between DNA and protein synthesis.
-Getting from DNA to protein requires two steps: transcription and translation.
-Transcription: synthesis of mRNA from a DNA template. Code is maintained due to complementary base pairing of RNA off DNA template ( A to T, U to A, C to G, G to C).
-occurs in the nucleus of eukaryotes
-Translation: synthesis of a polypeptide using the mRNA molecule at the ribosome.
-occurs in the cytoplasm of eukaryotes
-The basic mechanics of transcription and translation are similar for prokaryotes and eukaryotes.
- However, the lack of compartmentalization, due to the absence of a nucleus in prokaryotes, allows for translation to begin during transcription. In eukaryotes, the presence of a nucleus does not allows this.
-Eukaryotic mRNA is initially called a primary transcript ( or pre-mRNA) because it is modified prior to departing from the nucleus.
The Genetic Code
-There are only 4 nucleotide bases to specify 20 amino acids
Codons: Triplets of Nucleotides
-Triplets of nucleotide bases are the smallest units of uniform length that can code for all the amino acids.
-There are 64 possible codes (4(bases)^3(triplet))
-Triplet code: the genetic instructions for a polypeptide chain are written in the DNA as a series of non-overlapping, three nucleotide words.
- these "words" are then transcribed into corresponding non-overlapping, three nucleotide words in mRNA
-For each gene, only one of the DNA strands is transcribed, this is called the template strand
-mRNA molecules are complimentary and antiparallel to this template strand
-The corresponding compliment to a DNA triplet of mRNA is called a codon, and are synthesized in a 5' to 3' direction.
-during translation, codons are read in a 5' to 3' direction, each codon coding for a particular amino acid.
- the number of codons must be 3 times that of the amino acids which make up a polypeptide.
Cracking the Code
-In the 1960's each of the 64 RNA codon's were analyzed for coding
-Nirenberg: deciphered the first codon, UUU= phenylalanine
-Of the 64 codons, 3 code for "STOP" and 61 code for amino acids
-AUG, which codes for methionine, is often the start codon, though methionine may be removed later through modifications, newly synthesized polypeptides start with this amino acid.
-Multiple codons code for the same amino acid, but they usually only differ in the third nitrogenous base of the mRNA
-Reading Frame: nucleotides must be read in the correct non-overlapping groupings of 3 for correct translation
Evolution of the Genetic Code
-The code is nearly universal, shared by all organisms. All codons specify the same amino acids in every organism
-A language shared by all living things must have been operating very early in the history of life, early enough to be present in the common ancestor of all present-day organisms.
-The commonality allows us to utilize transformation to synthesize, for instance, human insulin in bacterial cells that have been transformed.
17.2: Transcription is the DNA-directed Synthesis of RNA
Molecular Components of Transcription
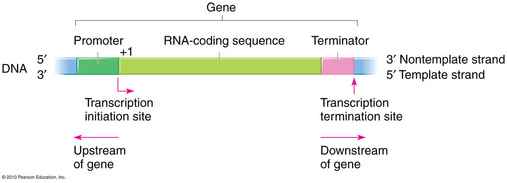
-Specific DNA sequences mark the beginning of the gene to be transcribed and the end.
-Promoter: the DNA sequence that initiates transcription (upstream from the transcription unit and terminator)
-Terminator: IN bacteria, the DNA sequence that signals the end of transcription (downstream from the promoter and transcription unit)
-Transcription unit: The segment of DNA that is transcribed (downstream from the promoter, upstream from the terminator)
-Not all proteins are enzymes, hypothesis reformatted to one gene-one protein
-Many proteins are composed of two or more polypeptide chains, hypothesis reformatted to one gene- one polypeptide
-Still not completely accurate since alternative splicing allows for a single gene to code for multiple polypeptides and some genes solely code for RNA and never become polypeptides.
Basic Principles of Transcription and Translation
-Genes code for proteins, but are not directly responsible for their synthesis. RNA bridges the gap between DNA and protein synthesis.
-Getting from DNA to protein requires two steps: transcription and translation.
-Transcription: synthesis of mRNA from a DNA template. Code is maintained due to complementary base pairing of RNA off DNA template ( A to T, U to A, C to G, G to C).
-occurs in the nucleus of eukaryotes
-Translation: synthesis of a polypeptide using the mRNA molecule at the ribosome.
-occurs in the cytoplasm of eukaryotes
-The basic mechanics of transcription and translation are similar for prokaryotes and eukaryotes.
- However, the lack of compartmentalization, due to the absence of a nucleus in prokaryotes, allows for translation to begin during transcription. In eukaryotes, the presence of a nucleus does not allows this.
-Eukaryotic mRNA is initially called a primary transcript ( or pre-mRNA) because it is modified prior to departing from the nucleus.
The Genetic Code
-There are only 4 nucleotide bases to specify 20 amino acids
Codons: Triplets of Nucleotides
-Triplets of nucleotide bases are the smallest units of uniform length that can code for all the amino acids.
-There are 64 possible codes (4(bases)^3(triplet))
-Triplet code: the genetic instructions for a polypeptide chain are written in the DNA as a series of non-overlapping, three nucleotide words.
- these "words" are then transcribed into corresponding non-overlapping, three nucleotide words in mRNA
-For each gene, only one of the DNA strands is transcribed, this is called the template strand
-mRNA molecules are complimentary and antiparallel to this template strand
-The corresponding compliment to a DNA triplet of mRNA is called a codon, and are synthesized in a 5' to 3' direction.
-during translation, codons are read in a 5' to 3' direction, each codon coding for a particular amino acid.
- the number of codons must be 3 times that of the amino acids which make up a polypeptide.
Cracking the Code
-In the 1960's each of the 64 RNA codon's were analyzed for coding
-Nirenberg: deciphered the first codon, UUU= phenylalanine
-Of the 64 codons, 3 code for "STOP" and 61 code for amino acids
-AUG, which codes for methionine, is often the start codon, though methionine may be removed later through modifications, newly synthesized polypeptides start with this amino acid.
-Multiple codons code for the same amino acid, but they usually only differ in the third nitrogenous base of the mRNA
-Reading Frame: nucleotides must be read in the correct non-overlapping groupings of 3 for correct translation
Evolution of the Genetic Code
-The code is nearly universal, shared by all organisms. All codons specify the same amino acids in every organism
-A language shared by all living things must have been operating very early in the history of life, early enough to be present in the common ancestor of all present-day organisms.
-The commonality allows us to utilize transformation to synthesize, for instance, human insulin in bacterial cells that have been transformed.
17.2: Transcription is the DNA-directed Synthesis of RNA
Molecular Components of Transcription
-Specific DNA sequences mark the beginning of the gene to be transcribed and the end.
-Promoter: the DNA sequence that initiates transcription (upstream from the transcription unit and terminator)
-Terminator: IN bacteria, the DNA sequence that signals the end of transcription (downstream from the promoter and transcription unit)
-Transcription unit: The segment of DNA that is transcribed (downstream from the promoter, upstream from the terminator)
-RNA polymerase: recognizes and binds the promoter sequence and pries DNA strands apart, joining nucleotides together, complimentary to the DNA template
RNA Polymerase vs DNA Polymerase:
-RNA Polymerase can only synthesize a nucleotide in a 5'-> 3' direction
-RNA Polymerase does not require primers to start the synthesis of a polynucleotide
Prokaryotic RNA polymerase vs Eukaryotic RNA polymerases:
-Bacteria have a single RNA polymerase that synthesizes all RNAs
-Eukaryotes have at least 3 RNA polymerases, RNA polymerase II synthesizes pre-mRNA, the others transcribe RNAs
that are not destined to become proteins.
Synthesis of and RNA Transcript
-There are 3 stages of Transcription: Initiation, elongation and termination.
RNA Polymerase Binding and Initiation of Transcript
-The promoter of a gene contains the start point, where synthesis begins.
-RNA Polymerase binds to the promoter in a specific location, identifying where transcription begins and which strand of DNA is the template.
-In bacteria, the RNA polymerase recognizes the promoter and binds to it alone.
-In eukaryotes, this binding is mediated by a collection of proteins called transcription factors.
-The transcription factors and RNA polymerase are referred to as the transcription initiation complex.
-These protein-protein interactions are important in controlling transcription, as transcription by RNA Pol. II cannot
bind the DNA and begin without all transcription factors bound to the template strand first.
-Eukaryotic promoters commonly include a TATA box, a sequence of DNA containing TATA, that is recognized by a transcription factor to begin the assembly of the transcription initiation complex.
Elongation of the RNA Strand
-RNA Pol. moves along DNA, untwisting the doubel helix to expose 10-20 nucleotides at a time, and adds nucleotides to the 3' end of the growing RNA molecule.
-As the RNA forms, it pulls away, allowing for the DNA double helix to reform.
-Several RNA polymerase molecules can transcribed the same gene simultaneously like trucks in a convoy following eachother.
-this increases the amount of mRNA that can be transcribed and helps the cell make the protein product in large amounts, if necessary.
Termination of Transcription
-Bacteria: transcription proceeds through the terminator sequence, and the synthesis of this sequence signals the end of transcription.
-RNA pol. is released, and the transcript is released.
-Bacterial transcripts are not modified, but complete at this point.
-Eukaryotes: RNA Pol. II transcribes a sequence on the DNA called the polyadenylation signal sequence.
-This results in a AAUAAA sequence on the pre-mRNA transcript, that is recognized and bound by proteins in the nucleus
-These proteins cut the pre-mRNA free from the polymerase, releasing it for further processing, prior to its release from the nucleus.
-RNA polymerase continues transcription until it is degraded by enzymes from its uncapped 5' end
17.3: Eukaryotic Cells Modify RNA after Transcription
Alteration of mRNA Ends
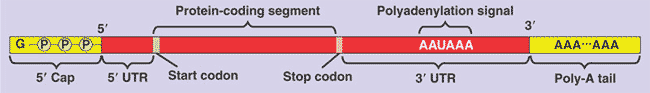
-A 5' cap, consisting of a modified guanine nucleotide, is added to the 5' end.
-A Poly-A tail, consisting of 50-250 Adenine nucleotides, is added to the 3' end following the polyadenylation signal.
-facilitate the export of the mRNA from the nucleus
-protect the mRNA from degradation by hydrolytic enzymes
-help ribosomes attach to 5' end of mRNA in the cytoplasm.
-In addition to the 5' cap and poly-A tail, there are two other untranslated regions (UTRs), one at the 5' end and one at the 3' end of
-These end modifications and UTRs will not be used during translation
RNA Polymerase vs DNA Polymerase:
-RNA Polymerase can only synthesize a nucleotide in a 5'-> 3' direction
-RNA Polymerase does not require primers to start the synthesis of a polynucleotide
Prokaryotic RNA polymerase vs Eukaryotic RNA polymerases:
-Bacteria have a single RNA polymerase that synthesizes all RNAs
-Eukaryotes have at least 3 RNA polymerases, RNA polymerase II synthesizes pre-mRNA, the others transcribe RNAs
that are not destined to become proteins.
Synthesis of and RNA Transcript
-There are 3 stages of Transcription: Initiation, elongation and termination.
RNA Polymerase Binding and Initiation of Transcript
-The promoter of a gene contains the start point, where synthesis begins.
-RNA Polymerase binds to the promoter in a specific location, identifying where transcription begins and which strand of DNA is the template.
-In bacteria, the RNA polymerase recognizes the promoter and binds to it alone.
-In eukaryotes, this binding is mediated by a collection of proteins called transcription factors.
-The transcription factors and RNA polymerase are referred to as the transcription initiation complex.
-These protein-protein interactions are important in controlling transcription, as transcription by RNA Pol. II cannot
bind the DNA and begin without all transcription factors bound to the template strand first.
-Eukaryotic promoters commonly include a TATA box, a sequence of DNA containing TATA, that is recognized by a transcription factor to begin the assembly of the transcription initiation complex.
Elongation of the RNA Strand
-RNA Pol. moves along DNA, untwisting the doubel helix to expose 10-20 nucleotides at a time, and adds nucleotides to the 3' end of the growing RNA molecule.
-As the RNA forms, it pulls away, allowing for the DNA double helix to reform.
-Several RNA polymerase molecules can transcribed the same gene simultaneously like trucks in a convoy following eachother.
-this increases the amount of mRNA that can be transcribed and helps the cell make the protein product in large amounts, if necessary.
Termination of Transcription
-Bacteria: transcription proceeds through the terminator sequence, and the synthesis of this sequence signals the end of transcription.
-RNA pol. is released, and the transcript is released.
-Bacterial transcripts are not modified, but complete at this point.
-Eukaryotes: RNA Pol. II transcribes a sequence on the DNA called the polyadenylation signal sequence.
-This results in a AAUAAA sequence on the pre-mRNA transcript, that is recognized and bound by proteins in the nucleus
-These proteins cut the pre-mRNA free from the polymerase, releasing it for further processing, prior to its release from the nucleus.
-RNA polymerase continues transcription until it is degraded by enzymes from its uncapped 5' end
17.3: Eukaryotic Cells Modify RNA after Transcription
Alteration of mRNA Ends
-A 5' cap, consisting of a modified guanine nucleotide, is added to the 5' end.
-A Poly-A tail, consisting of 50-250 Adenine nucleotides, is added to the 3' end following the polyadenylation signal.
-facilitate the export of the mRNA from the nucleus
-protect the mRNA from degradation by hydrolytic enzymes
-help ribosomes attach to 5' end of mRNA in the cytoplasm.
-In addition to the 5' cap and poly-A tail, there are two other untranslated regions (UTRs), one at the 5' end and one at the 3' end of
-These end modifications and UTRs will not be used during translation
Split Genes and RNA Splicin'
RNA Splicing: the removal of large portions of the RNA molecule that is initially synthesized.
-Intron (intervening sequence): noncoding segment (not translated) that lies between coding segments of DNA strands and their RNA compliments.
-Introns are removed during the splicing process by a large complex of proteins and RNAs called a spliceosome.
-Exon (expressed sequence): coding segment (translated to a protein) of DNA strand and their RNA compliments
- once introns are removed adjacent exons are joined by the same spliceosome complex.
Ribozymes
-Ribozymes: catalytic RNAs that participate as enzymes in the spliceosome.
-In some organisms, the introns act as their own ribozymes, excising themselves.
-Therefore, all catalytic enzymes are not proteins.
-Three properties enable RNA to act as enzymes:
1) The single-strandedness of RNA allows it to self adhere to complimentary segments elsewhere in the molecule, allowing
RNAs to take the 3D shape necessary for specific binding properties like a protein.
2) Some bases in RNAs contain functional groups (like the amino acids of a protein) that can participate in catalytic reactions.
3) RNA can hydrogen bond with other RNA molecules or DNA, adding specificity of binding.
The Functional and Evolutionary Importance of Introns
-Some introns have been identified as containing sequences that regulate gene expression and many affect gene products.
-Many genes code for multiple proteins, depending on which segments are removed as non-coding introns, this is called alternative splicing.
- because of this the number of protein products an organism produces can be greater than the amount of genes it contains.
-domain: discrete structural and functional regions of a protein.
- In multiple cases, the different exons code for specific domains.
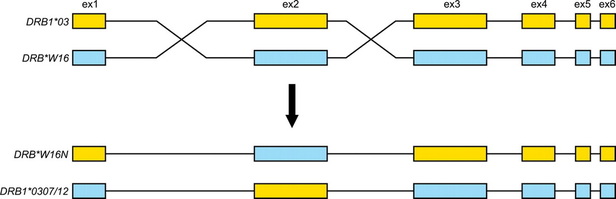
-The presence of introns may facilitate the evolution of new and potentially beneficial proteins via exon shuffling. (see below, introns are black lines between colored exons)
-introns increase the probability of crossing over between the exons of alleles of a gene by providing more terrain for cross overs without interrupting coding sequences.
- this could result in new combos of exons and new protein products with altered structures, and therefore, altered functions.
RNA Splicing: the removal of large portions of the RNA molecule that is initially synthesized.
-Intron (intervening sequence): noncoding segment (not translated) that lies between coding segments of DNA strands and their RNA compliments.
-Introns are removed during the splicing process by a large complex of proteins and RNAs called a spliceosome.
-Exon (expressed sequence): coding segment (translated to a protein) of DNA strand and their RNA compliments
- once introns are removed adjacent exons are joined by the same spliceosome complex.
Ribozymes
-Ribozymes: catalytic RNAs that participate as enzymes in the spliceosome.
-In some organisms, the introns act as their own ribozymes, excising themselves.
-Therefore, all catalytic enzymes are not proteins.
-Three properties enable RNA to act as enzymes:
1) The single-strandedness of RNA allows it to self adhere to complimentary segments elsewhere in the molecule, allowing
RNAs to take the 3D shape necessary for specific binding properties like a protein.
2) Some bases in RNAs contain functional groups (like the amino acids of a protein) that can participate in catalytic reactions.
3) RNA can hydrogen bond with other RNA molecules or DNA, adding specificity of binding.
The Functional and Evolutionary Importance of Introns
-Some introns have been identified as containing sequences that regulate gene expression and many affect gene products.
-Many genes code for multiple proteins, depending on which segments are removed as non-coding introns, this is called alternative splicing.
- because of this the number of protein products an organism produces can be greater than the amount of genes it contains.
-domain: discrete structural and functional regions of a protein.
- In multiple cases, the different exons code for specific domains.
-The presence of introns may facilitate the evolution of new and potentially beneficial proteins via exon shuffling. (see below, introns are black lines between colored exons)
-introns increase the probability of crossing over between the exons of alleles of a gene by providing more terrain for cross overs without interrupting coding sequences.
- this could result in new combos of exons and new protein products with altered structures, and therefore, altered functions.
17.4: Translation is the RNA-directed Synthesis of a Polypeptide
Molecular Components of Translation
-transfer RNA (tRNA): transfers amino acids from the cytoplasm to a growing polypeptide in the ribosome by translating the mRNA codons into amino acids.
The Structure and Function of Transfer RNA
-Each tRNA molecule is single-stranded and consists of about 80 nucleotides, which can base-pair with other complimentary segments of the same molecule, giving the tRNA a 3-D shape that is rougly L-shaped
-one end bears a specific amino acid bound to the 3' end
- the loop of tRNA extending from the bottom of the tRNA molecule contains a nucleotide triplet, called an anticodon, that can base pair with the complimentary codon on mRNA.
-The tRNA "reads" the codons on the mRNA, one at a time, and delivers the correct amino acid, based on the anticodon that compliments the mRNA codon.
-tRNA molecules are synthesized in the nucleus from DNA templates, and travel to the cytosol to be used repetitively in translation.
-Once they deliver their specified amino acid, they leave the ribosome to fetch another amino acid of the same identity,
waiting to be called again.
-The correct matching of tRNA and amino acid is carried out by enzymes called aminoacyl-tRNA synthetases.
-there are 20 of these synthetases, one for each amino acid.
-Some tRNAs can bind multiple codons, because there are several codons that code for the same amino acids.
- for this reason, complimentary binding of a tRNA anticodon to a codon is relaxed at the 3rd base of the codon.
-This flexible pairing at the 3rd nucleotide of an mRNA codon is called wobble.
Ribosomes
-Ribosomes facilitate the coupling of tRNA anticodons and mRNA codons during protein synthesis.
-Ribosomes are made up of two subunits, a large subunit and a small subunit composed of ribosomal RNA (rRNA) and proteins
-rRNAs are made in the nucleolus of eukaryotic cells and exported to the cytoplasm.
-rRNA is the most abundant type of RNA in a cell, due to an overwhelming presence of ribosomes.
-rRNA is primarily responsible for the function of the ribosome, the protein components merely appear to support the shaping
of the rRNA.
-Ribosomal subunits join to form a functional ribosome only when attached to an mRNA molecule:
-Structure of a bacterial ribosome:
-P site: holds tRNA carrying the growing polypeptide chain
-A site: holds tRNA carrying the next amino acid to be added
-E site: discharged tRNas exit the ribosome.
-When a polypeptide is complete it exits through a tunnel in the large subunit.
Building a Polypeptide
-Translation is also divided into initiation, elongation and termination.
- all three stages require protein factors that aid in the translation process
- it also requires energy that is provided by GTP
Ribosome Association and Initiation of Translation
-The small subunit of the ribosome binds mRNA at a specific RNA sequence upstream from the start codon (AUG in eukaryotes) and the specific initiator tRNA, which carries the start amino acid, methionine.
- The start codon signals the beginning of translation and establishes the reading frame.
-This is followed by the attachment of the large ribosomal subunit, which completes the initiation complex.
-Proteins called initiation factors are required bring all the components together, as is the hydrolysis of a GTP molecule.
- once the complex is complete the initiator tRNA sits in the P site, leaving the A site open for the next tRNA
- amino acids are added to the carboxyl end of the growing polypeptide chain
Elongation of the Polypeptide Chain
-amino acids are added one by one to the previous amino acid's carboxyl end
-each addition requires proteins called elongation factors.
1) Codon recognition: the appropriate tRNA base pairs with complimentary mRNA codon in the A site.
2) Peptide bond formation: rRNA molecule of large subunit catalyzes formation of peptide bond between amino group of new amino acid and carboxyl group of existing amino acid, attaching the growing polypeptide chain to the new tRNA molecule.
3) Translocation: the tRNA in the A site, with the polypeptide chain now attached is moved into the P site and the tRNA that was in the P site is moved to the E site. The mRNA moves along, bringing the next codon into the A site.
Termination of Translation
-Elongation continues until a stop codon reaches the A site.
-Stop codons (UAG, UAA, and UGA) do not code for amino acids, but stop translation.
-The stop codon is bound by release factor protein, which adds a water molecule to the polypeptide chain instead of an amino acid, hydrolyzing the bond between the tRNA and polypeptide chain in the P site, and releasing the polypeptide through the exit tunnel of the large ribosomal subunit.
-The translation complex is then broken down.
Completing and Targeting the Functional Protein
-translation alone does not make a functional protein, polypeptides undergo modifications
Protein Folding and Post-Translational Modifications
-During synthesis polypeptides begin to coil and fold spontaneously (primary structure: amino acid sequence), forming a protein with a specific 3D shape with secondary (backbone interactions) and tertiary (side chain interactions) structures.
- A gene determines primary structure and primary structure determines secondary and tertiary structures, and therefore the protein's shape.
- a protein called a chaperonin may aid in protein folding by sheltering it from the cytosolic environment.
-Proteins may undergo post-translational modifications: amino acids may be chemically modified by the addition of other molecules, amino acids may be removed from the amino end of the polypeptide, the polypeptide may be cleaved into two separate protein products, or two or more polypeptides may come together to form a protein with quarternary structure ( 2 or more polypeptides interacting).
Targeting Polypeptides to Specific Locations
There are 2 types of ribosomes that are identical in structure and can alternate between types:
1) Free ribosomes: located in cytosol, synthesize proteins that function in the cytosol.
2) Bound ribosomes: attached to cytosolic side of rough ER or to the nuclear envelope, syntehsize proteins of the endomembrane system (nuclear envelope, ER, golgi, lysosomes, vacuoles, plasma membrane) and well as proteins for secretion from the cell.
All protein synthesis begins in the cytosol, however, polypeptides of proteins destined for the endomembrane system are marked by a signal peptide.
-This signal peptide is recognized as it emerged from the ribosome by a protein/rRNA complex called a signal recognition particle (SRP), that escorts it to the ER membrane for translocation , where translation continues, and the peptide elongates into the ER lumen
Proteins that have destinations outside of the endomembrane system are tagged differently, but they are completely synthesized in the cytosol, prior to relocation.
Making Multiple Polypeptides in Bacteria and Eukaryotes
-When a polypeptide is required by the cell multiple ribosomes can translate the same mRNA molecule and make many copies of the same polypeptide simultaneously, allowing for quick production of the protein.
- when a string of ribosomes all bind the same mRNA molecule, it is called a polyribosome or polysome.
-Another way that quick protein production can be established is by transcribing multiple mRNA molecules, however this process is more efficient in bacteria because compartmentalization and modifications of eukaryotic mRNA slow the release of the mRNA from the nucleus.
17.5: Mutations of One or a Few Nucleotides can Affect Protein Structure
-Mutation: changes to the genetic information of a cell, responsible for the diversity of genes among organisms because they are the source of new genes.
-Point mutation: changes in a single nucleotide pair of a gene.
- only passed to offspring if it occurs in a gamete or in a cell that produces gametes.
- when a mutation is passed to offspring, resulting in an unfavorable phenotype, it is called a genetic disorder or hereditary disease. (i.e: sickle cell anemia, familial cardiomyopathy)
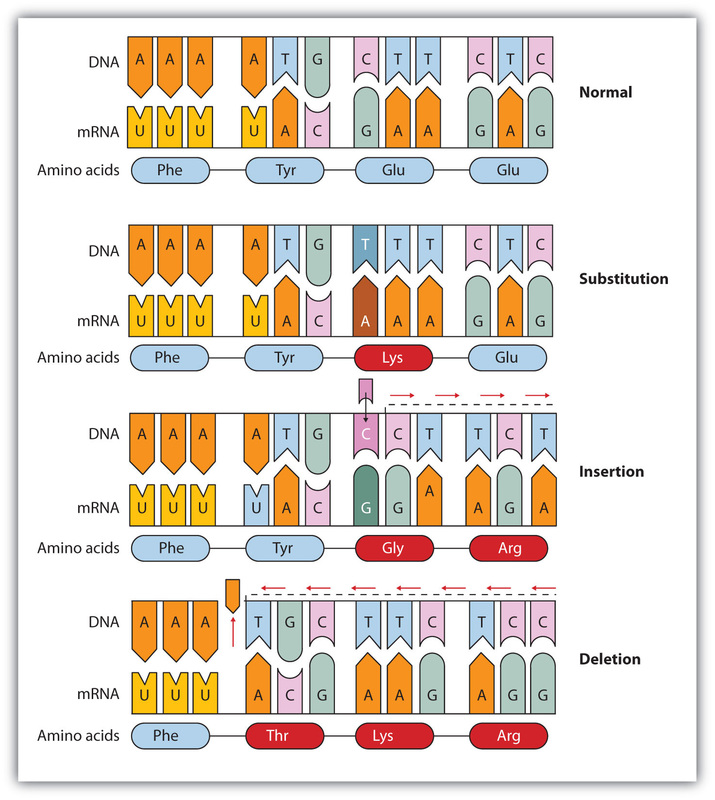
Types of Small-Scale Mutations
-Two types of point mutations: single nucleotide pair substitutions and nucleotide pair insertions or deletions.
Substitutions
-nucleotide-pair substitution: replacement of one nucleotide and its partner with another pair of nucleotides.
1) substitution may have no effect because several codons code for the same amino acids, when this occurs it is called a silent mutation.
2) substitutions that cause the codon to code for a different amino acid are called missense mutations, and can result in one of three outcomes:
- this may not have a significant affect on the protein product depending on the location of the incorrect amino acid in the protein's structure and whether or not the incorrect amino acid has properties similar to that of the correct amino acid.
- it could have a significant affect if the incorrect amino acid is located at a point that is crucial to the structure and function of the protein (like in sickle cell).
-the incorrect protein could be an improved version of the intended protein, though most tend to be neutral or detrimental.
3) substitutions that change a codon for an amino acid to a stop codon are called nonsense mutations. These result in the early termination of translation, and therefore, an incomplete polypeptide, which most often is nonfunctional.
Insertions and Deletions
-insertions: addition of nucleotide pairs in a gene
-deletions: losses of nucleotide pairs in a gene.
-These mutations tend to be disastrous more often than substitutions because they can alter the reading frame permanently, while substitutions solely affect the interpretation of a single codon.
-frameshift mutation: whenever the number of nucleotides inserted or deleted is not a multiple of three. This results in the reading frame being permanently altered downstream from the mutation, making for multiple missense occurrences, usually
resulting in nonsense at some point, and premature termination.
- unless this occurs at the very end of a polypeptide, it is unlikely to be functional.
Molecular Components of Translation
-transfer RNA (tRNA): transfers amino acids from the cytoplasm to a growing polypeptide in the ribosome by translating the mRNA codons into amino acids.
The Structure and Function of Transfer RNA
-Each tRNA molecule is single-stranded and consists of about 80 nucleotides, which can base-pair with other complimentary segments of the same molecule, giving the tRNA a 3-D shape that is rougly L-shaped
-one end bears a specific amino acid bound to the 3' end
- the loop of tRNA extending from the bottom of the tRNA molecule contains a nucleotide triplet, called an anticodon, that can base pair with the complimentary codon on mRNA.
-The tRNA "reads" the codons on the mRNA, one at a time, and delivers the correct amino acid, based on the anticodon that compliments the mRNA codon.
-tRNA molecules are synthesized in the nucleus from DNA templates, and travel to the cytosol to be used repetitively in translation.
-Once they deliver their specified amino acid, they leave the ribosome to fetch another amino acid of the same identity,
waiting to be called again.
-The correct matching of tRNA and amino acid is carried out by enzymes called aminoacyl-tRNA synthetases.
-there are 20 of these synthetases, one for each amino acid.
-Some tRNAs can bind multiple codons, because there are several codons that code for the same amino acids.
- for this reason, complimentary binding of a tRNA anticodon to a codon is relaxed at the 3rd base of the codon.
-This flexible pairing at the 3rd nucleotide of an mRNA codon is called wobble.
Ribosomes
-Ribosomes facilitate the coupling of tRNA anticodons and mRNA codons during protein synthesis.
-Ribosomes are made up of two subunits, a large subunit and a small subunit composed of ribosomal RNA (rRNA) and proteins
-rRNAs are made in the nucleolus of eukaryotic cells and exported to the cytoplasm.
-rRNA is the most abundant type of RNA in a cell, due to an overwhelming presence of ribosomes.
-rRNA is primarily responsible for the function of the ribosome, the protein components merely appear to support the shaping
of the rRNA.
-Ribosomal subunits join to form a functional ribosome only when attached to an mRNA molecule:
-Structure of a bacterial ribosome:
-P site: holds tRNA carrying the growing polypeptide chain
-A site: holds tRNA carrying the next amino acid to be added
-E site: discharged tRNas exit the ribosome.
-When a polypeptide is complete it exits through a tunnel in the large subunit.
Building a Polypeptide
-Translation is also divided into initiation, elongation and termination.
- all three stages require protein factors that aid in the translation process
- it also requires energy that is provided by GTP
Ribosome Association and Initiation of Translation
-The small subunit of the ribosome binds mRNA at a specific RNA sequence upstream from the start codon (AUG in eukaryotes) and the specific initiator tRNA, which carries the start amino acid, methionine.
- The start codon signals the beginning of translation and establishes the reading frame.
-This is followed by the attachment of the large ribosomal subunit, which completes the initiation complex.
-Proteins called initiation factors are required bring all the components together, as is the hydrolysis of a GTP molecule.
- once the complex is complete the initiator tRNA sits in the P site, leaving the A site open for the next tRNA
- amino acids are added to the carboxyl end of the growing polypeptide chain
Elongation of the Polypeptide Chain
-amino acids are added one by one to the previous amino acid's carboxyl end
-each addition requires proteins called elongation factors.
1) Codon recognition: the appropriate tRNA base pairs with complimentary mRNA codon in the A site.
2) Peptide bond formation: rRNA molecule of large subunit catalyzes formation of peptide bond between amino group of new amino acid and carboxyl group of existing amino acid, attaching the growing polypeptide chain to the new tRNA molecule.
3) Translocation: the tRNA in the A site, with the polypeptide chain now attached is moved into the P site and the tRNA that was in the P site is moved to the E site. The mRNA moves along, bringing the next codon into the A site.
Termination of Translation
-Elongation continues until a stop codon reaches the A site.
-Stop codons (UAG, UAA, and UGA) do not code for amino acids, but stop translation.
-The stop codon is bound by release factor protein, which adds a water molecule to the polypeptide chain instead of an amino acid, hydrolyzing the bond between the tRNA and polypeptide chain in the P site, and releasing the polypeptide through the exit tunnel of the large ribosomal subunit.
-The translation complex is then broken down.
Completing and Targeting the Functional Protein
-translation alone does not make a functional protein, polypeptides undergo modifications
Protein Folding and Post-Translational Modifications
-During synthesis polypeptides begin to coil and fold spontaneously (primary structure: amino acid sequence), forming a protein with a specific 3D shape with secondary (backbone interactions) and tertiary (side chain interactions) structures.
- A gene determines primary structure and primary structure determines secondary and tertiary structures, and therefore the protein's shape.
- a protein called a chaperonin may aid in protein folding by sheltering it from the cytosolic environment.
-Proteins may undergo post-translational modifications: amino acids may be chemically modified by the addition of other molecules, amino acids may be removed from the amino end of the polypeptide, the polypeptide may be cleaved into two separate protein products, or two or more polypeptides may come together to form a protein with quarternary structure ( 2 or more polypeptides interacting).
Targeting Polypeptides to Specific Locations
There are 2 types of ribosomes that are identical in structure and can alternate between types:
1) Free ribosomes: located in cytosol, synthesize proteins that function in the cytosol.
2) Bound ribosomes: attached to cytosolic side of rough ER or to the nuclear envelope, syntehsize proteins of the endomembrane system (nuclear envelope, ER, golgi, lysosomes, vacuoles, plasma membrane) and well as proteins for secretion from the cell.
All protein synthesis begins in the cytosol, however, polypeptides of proteins destined for the endomembrane system are marked by a signal peptide.
-This signal peptide is recognized as it emerged from the ribosome by a protein/rRNA complex called a signal recognition particle (SRP), that escorts it to the ER membrane for translocation , where translation continues, and the peptide elongates into the ER lumen
Proteins that have destinations outside of the endomembrane system are tagged differently, but they are completely synthesized in the cytosol, prior to relocation.
Making Multiple Polypeptides in Bacteria and Eukaryotes
-When a polypeptide is required by the cell multiple ribosomes can translate the same mRNA molecule and make many copies of the same polypeptide simultaneously, allowing for quick production of the protein.
- when a string of ribosomes all bind the same mRNA molecule, it is called a polyribosome or polysome.
-Another way that quick protein production can be established is by transcribing multiple mRNA molecules, however this process is more efficient in bacteria because compartmentalization and modifications of eukaryotic mRNA slow the release of the mRNA from the nucleus.
17.5: Mutations of One or a Few Nucleotides can Affect Protein Structure
-Mutation: changes to the genetic information of a cell, responsible for the diversity of genes among organisms because they are the source of new genes.
-Point mutation: changes in a single nucleotide pair of a gene.
- only passed to offspring if it occurs in a gamete or in a cell that produces gametes.
- when a mutation is passed to offspring, resulting in an unfavorable phenotype, it is called a genetic disorder or hereditary disease. (i.e: sickle cell anemia, familial cardiomyopathy)
Types of Small-Scale Mutations
-Two types of point mutations: single nucleotide pair substitutions and nucleotide pair insertions or deletions.
Substitutions
-nucleotide-pair substitution: replacement of one nucleotide and its partner with another pair of nucleotides.
1) substitution may have no effect because several codons code for the same amino acids, when this occurs it is called a silent mutation.
2) substitutions that cause the codon to code for a different amino acid are called missense mutations, and can result in one of three outcomes:
- this may not have a significant affect on the protein product depending on the location of the incorrect amino acid in the protein's structure and whether or not the incorrect amino acid has properties similar to that of the correct amino acid.
- it could have a significant affect if the incorrect amino acid is located at a point that is crucial to the structure and function of the protein (like in sickle cell).
-the incorrect protein could be an improved version of the intended protein, though most tend to be neutral or detrimental.
3) substitutions that change a codon for an amino acid to a stop codon are called nonsense mutations. These result in the early termination of translation, and therefore, an incomplete polypeptide, which most often is nonfunctional.
Insertions and Deletions
-insertions: addition of nucleotide pairs in a gene
-deletions: losses of nucleotide pairs in a gene.
-These mutations tend to be disastrous more often than substitutions because they can alter the reading frame permanently, while substitutions solely affect the interpretation of a single codon.
-frameshift mutation: whenever the number of nucleotides inserted or deleted is not a multiple of three. This results in the reading frame being permanently altered downstream from the mutation, making for multiple missense occurrences, usually
resulting in nonsense at some point, and premature termination.
- unless this occurs at the very end of a polypeptide, it is unlikely to be functional.
New Mutations and Mutagens
-errors during replication or recombination can lead to these mutations, these are called spontaneous mutations when they elude the proofreading machinery.
-mutagen: physical or chemical agent that interact with DNA to cause mutations
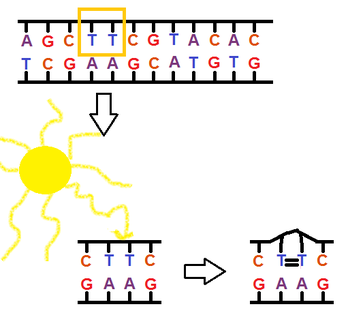
-x-rays and other forms of high-energy radiation pose hazards to the genetic material of the exposed organism
- UV-light falls in this category of mutagenic radiation, causing thymine dimers.
-errors during replication or recombination can lead to these mutations, these are called spontaneous mutations when they elude the proofreading machinery.
-mutagen: physical or chemical agent that interact with DNA to cause mutations
-x-rays and other forms of high-energy radiation pose hazards to the genetic material of the exposed organism
- UV-light falls in this category of mutagenic radiation, causing thymine dimers.
-Chemical mutagens:
-nucleotide analogs are chemicals similar to normal DNA nucleotides, but pair incorrectly during DNA replication.
-others interfere with correct DNA replication by inserting themselves into the DNA, distorting the double helix.
-other cause chemical changes to the bases, that change their pairing properties.
-Carcinogen: cancer-causing chemicals, most are mutagenic (and most mutagens are carcinogenic)
-nucleotide analogs are chemicals similar to normal DNA nucleotides, but pair incorrectly during DNA replication.
-others interfere with correct DNA replication by inserting themselves into the DNA, distorting the double helix.
-other cause chemical changes to the bases, that change their pairing properties.
-Carcinogen: cancer-causing chemicals, most are mutagenic (and most mutagens are carcinogenic)